About CHILDdb
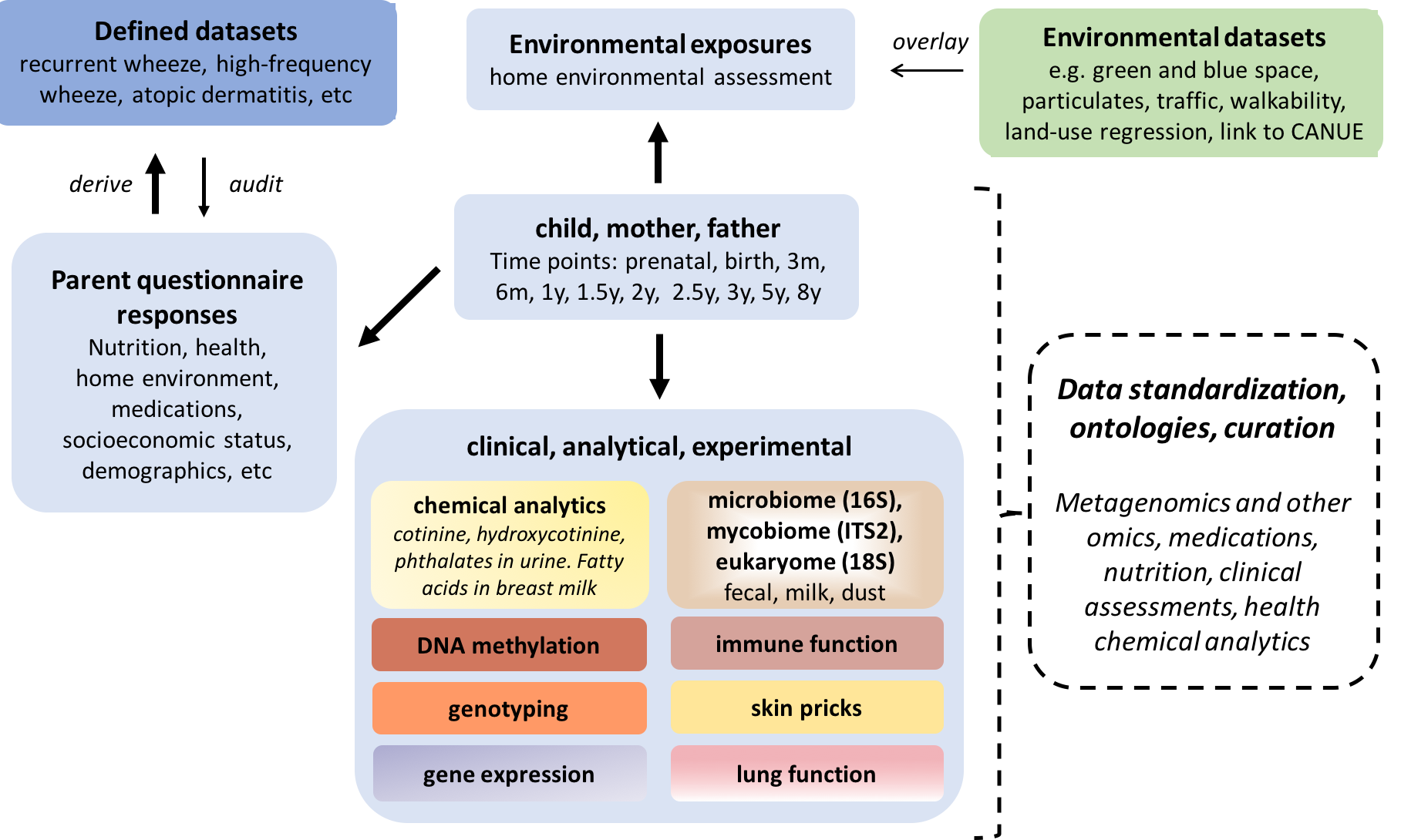
A centralized database integrating CHILD cohort data (e.g. health, nutrition, environmental, socioeconomic and other factors) with external analytic/experimental data (e.g. microbiome, epigenetics, metabolomics) is critical for enabling more integrative analyses of diverse data generated by the CHILD study. CHILDdb is critical to fully capitalizing on our rich data through integraton of these diverse datasets.
CHILDdb utilizes standardized, hierarchical terms called ontologies for describing select data. Ontologies help overcome many of the challenges of querying free-text data describing contextual information – especially when they are derived from different sources. For example, to improve hierarchical querying of medications, free-text medications in the database have been curated, standardized and mapped to 620 controlled vocabulary terms representing generic medication names and nutraceuticals from the Chemical Entities of Biological Interest (ChEBI) ontology (https://www.ebi.ac.uk/chebi/; an ontology containing hierarchical relationships between various drugs and their classes, as well as NCIT, the Drug Ontology and DrugBank (drugbank.ca). This enables queries like comparing subjects’ who had taken any non-steroidal anti-inflammatory versus those that had not, by a given time point (or identifying differences, such as microbiome differences, associated with certain drug class use). We have identified other CHILD data that can benefit from similar integration of other ontologies – enabling more complex, hierarchical analyses. These ontologies are also key facilitators allowing harmonization of CHILD data with other cohorts.
Current development and production versions of CHILDdb are hosted and administered by the SFU Research Computing Group (RCG).
Overview of CHILDdb
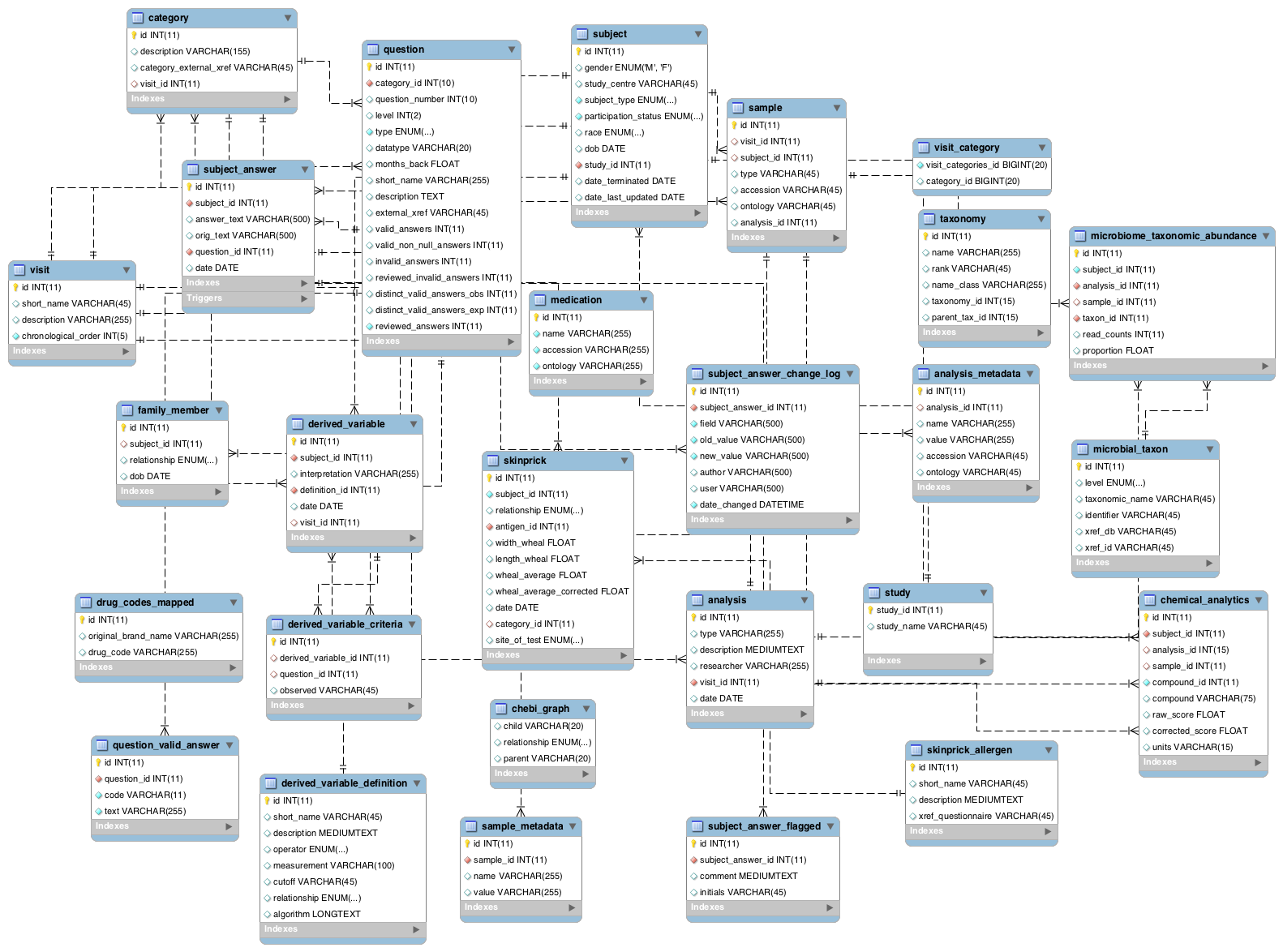
Overview of Core Components of MySQL Schema Developed for CHILDdb